1 微生物组数据使用TMM等矫正测序数据-更加适合目前发文章要求
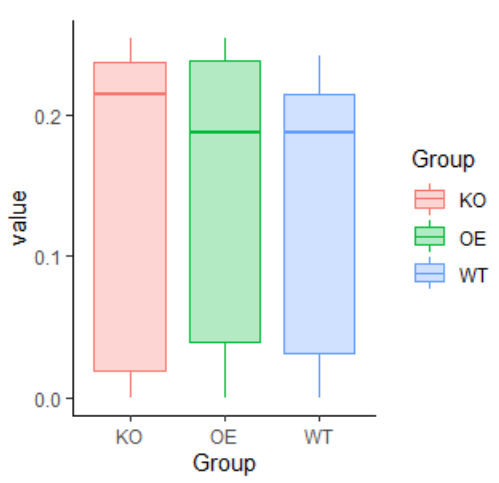
2 使用箱线图可视化距离矩阵
library(tidyverse)library(ggClusterNet)library(phyloseq)data(ps)ps.scale = scale_micro(ps,method = "TMM" )unif <- phyloseq::distance(ps.scale , method= "bray") %>%as.matrix()gro = sample_data(ps)head(gro)library(tidyfst)dat1 = unif %>%as.matrix() %>%tidyfst::mat_df()head(dat1)A = c()for (i in 1:dim(dat1)[1]) {if (gro[dat1$row[i],][[2]] == gro[dat1$col[i],][[2]]) {A[i] = T} else {A[i] =F}}dat1$group = Ahead(gro)gro$id = row.names(gro)dat2 = dat1 %>% filter(group == T) %>%left_join(gro,by = c("row" = "id"))head(dat2)ggplot(dat2) +geom_boxplot(aes(x = Group,y = value,color =Group,fill = Group),alpha = 0.3) +theme_classic()